Development News Brief
new: % hg clone http://www.bx.psu.edu/hg/galaxy galaxy-dist
upgrade: % hg pull -u -r ca0c4ad2bb39
What's New
galaxyproject at Twitter
Join us at Twitter for the latest updates on development projects, conferences and training, and all things Galaxy!
#usegalaxy
http://twitter.com/#!/search/galaxyproject
New Ways to Search Data Libraries
In addition to searching on data library names and descriptions, we have introduced the ability to search on attributes of library datasets.
Two different approaches to searching library datasets are supported
The public Galaxy instances hosted at Penn State University use the whoosh implementation, while Brad Chapman's instance uses the * Lucene * implementation. Choose either of these implementations for your local Galaxy instances (but not both).
There are two new config settings in the "Data Libraries" section of universe_wsgi.ini.sample for these implementations (again, uncomment only one of these if you choose to use this feature):
# Search data libraries with whoosh
#enable_whoosh_library_search = True
# Whoosh indexes are stored in this directory.
#whoosh_index_dir = database/whoosh_indexes
# Search data libraries with lucene
#enable_lucene_library_search = False
# maxiumum file size to index for searching, in MB
#fulltext_max_size = 500
#fulltext_noindex_filetypes=bam,sam,wig,bigwig,fasta,fastq,fastqsolexa,fastqillumina,fastqsanger
# base URL of server providing search functionality using lucene
#fulltext_url = http://localhost:8081
Details for searching with whoosh
You will need to build indexes on a regular, timely basis for searching your library datasets. You can do this by running the following script included in the distribution:
% <your galaxy install dir>/scripts/data_libraries/build_whoosh_index.sh
Running this script will build whoosh indexes in the directory named by the config setting labeled whoosh_index_dir shown above.
We recommend adding something like the following setting to your Galaxy server's cron settings to keep the indexes current:
0,30 * * * * cd /var/opt/galaxy/g2test/galaxy_test ; bash ./scripts/data_libraries/build_whoosh_index.sh
The above setting rebuilds the indexes every 30 minutes. You may want to build them more often because search results will include only those library datasets that were indexed the last time the script was executed. In other words, if a library dataset was uploaded after the whoosh indexes were created, it will not be returned in search results even if it meets the search criteria.
Using whoosh, you can search on the "name", "info", "message" and "dbkey" attributes of library datasets using the following new search box displayed at the top of the "Data Libraries" page:
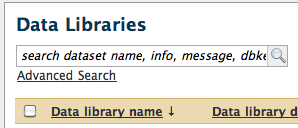
Clicking the "Advanced search" link below the search box allows you to search data library names and descriptions as you did in the past:
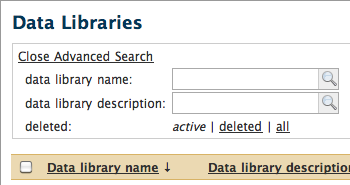
Details for searching with Lucene
If you use the Lucene implementation instead of whoosh, there are a few differences. You'll need to build Lucene indexes using the following script:
<your galaxy install dir>/scripts/data_libraries/build_lucene_index.sh
In addition to the "name", "info", "message" and "dbkey" attributes, this implementation will also index the contents of your library datasets. You should decide on the optimal values for the following config settings for your Galaxy instance:
# maxiumum file size to index for searching, in MB
#fulltext_max_size = 500
#fulltext_noindex_filetypes=bam,sam,wig,bigwig,fasta,fastq,fastqsolexa,fastqillumina,fastqsanger
You'll also need to decide on a value for the following config setting:
# base URL of server providing search functionality using lucene
#fulltext_url = http://localhost:8081
Managing disk space with Data Libraries
Adjusted the cleanup_datasets.py script to more correctly handle the lifecycle of Library Datasets.
Managing library datasets is a bit complex, so here is a scenario that hopefully provides clarification. The complexities of handling library datasets is mostly contained in the delete_datasets() method in the cleanup_datasets.py script.
Example of usage:
- Assume we have 1 library dataset with
LibraryDatasetDatasetAssociation -> LibraryDataset and Dataset
This dataset would have the following database column values:
LibraryDatasetDatasetAssociation deleted: False
LibraryDataset deleted: False, purged: False
Dataset deleted: False purged: False
Now a user deletes the associated dataset from a data library via a UI menu option.
This action results in the following database column values:
LibraryDatasetDatasetAssociation deleted: False
LibraryDataset deleted: True*, purged: False
Dataset deleted: False, purged: False
- After the number of days configured for the
delete_datasets()method (option -6 in thecleanup_datasets.pyscript) have passed, execution of thedelete_datasets()method results in the following database column values:
LibraryDatasetDatasetAssociation deleted: True
LibraryDataset deleted: True, purged: True
Dataset deleted: True*, purged: False
- After the number of days configured for the
purge_datasets()method (option -3 in thecleanup_datasets.pyscript) have passed, execution of thepurge_datasets()method results in the following database column values:
LibraryDatasetDatasetAssociation deleted: True
LibraryDataset deleted: True, purged: True
Dataset deleted: True, purged: True
(dataset file removed from disk if -r flag is used)
This scenario is about as simple as it gets. Keep in mind that a Dataset object can have many HistoryDatasetAssociations and many LibraryDatasetDatasetAssociations, and a LibraryDataset can have many LibraryDatasetDatasetAssociations.
Another way of stating it is: LibraryDatasetDatasetAssociation objects map LibraryDataset objects to Dataset objects, and Dataset objects may be mapped to History objects via HistoryDatasetAssociation
Updated & Improved
Current Tools
- Update FASTX tool wrappers to handle
fastqsangerformats. - The BLAST+ tools are now uncommented in the
sample tool configuration. - Fix an
Rpysyntax problem in theHistogramtool that was causing the tool to fail in some local installations. - Enable Cuffcompare to take an arbitrary number of input
GTFfiles. - Add
GTF sniffertodatatypes configsample file.
New Tools
- Added a Line/Word/Character counter to
Text Manipulationtool group. -
Extended
Extract Genomic DNAtool to support GFF/GTF features and custom genomes:- This is especially useful for extracting genomic data that correspond to transcripts in GTF format, as is produced by
Cufflinks. - Tool also now accepts sequence data from a history item, enabling the extraction of data from custom genomes.
- This is especially useful for extracting genomic data that correspond to transcripts in GTF format, as is produced by
New Community Tools Added (Tool Shed)
- Velvet, ABySS, Minimus2, Phrap, and Newbler. http://community.g2.bx.psu.edu/
Data Libraries
- Fix a bug that included the option for a regular (non-admin) user to upload a directory of files when the
configsetting existed, but the user's directory did not exist. Instead of throwing an exception, an attempt to create the directory will now be made. If the attempt fails, the option to "Upload a directory of files" will not be included in the select list on the upload form for that particular user. - When browsing a data library, only display the check boxes and the actions to perform on multiple selected datasets if library datasets are displayed.
- Clean up the behavior when performing an action on multiple library datasets. Instead of throwing an error and not allowing the action on any of the datasets if the user is not authorized to perform the action on one of them, all "unauthorized" datasets are now left alone and the action is performed on all datasets for which the user is authorized to perform the action. Appropriate messages detail what occurred.
- Allow regular users to delete multiple datasets returned from library dataset searches based on the behavior described above.
- Do not include the
"Select datasets for import into selected histories"option in the actions popup menu at the data library level if the library's root folder doesn't contain any accessible datasets. Make behavior the same for folder popup menus. - Fix a bug where a regular user could never see deleted library items when toggling show / hide deleted items in a data library.
- Add a purged column to the
LibraryDatasettable. ALibraryDatasetis marked "purged" when all associatedLibraryDatasetDatasetAssociationsare marked "deleted".
Workflows
- Workflow steps on the
"Run workflow"page are now collapsible to just the title bar with the tool name and annotation. Steps without any inputs to be set will be collapsed by default. - When running a workflow, possible values for
Input Datasetsteps are filtered todatatypesthat are valid for immediately subsequent steps, instead of just using 'data'. - Jquery1.5 workflow fixes with quotations and flagging of outputs.
Workflows containing tools that are not loaded will now open in the editor instead of failing on the dictionary lookup. The offending nodes are marked as having an error state, and the workflow cannot be saved. This will enable users to at least look at and try to recover a workflow by removing (and potentially replacing) the offending steps, as well as providing a better view of what's actually wrong with the workflow.
- Workflows will no longer try to run when required tools are not found. An error message is displayed with a link to the editor view (to fix the workflow).
- Interval datatype now uses line estimates for large files. This fixes potential "'?' lines" seen by some users.
- Removed "Annotation:" labels for a cleaner look on the run workflow page.
- New History functionality now shows only the checkbox by default, displaying the input for a history name only when checked.
- Tool text has been standardized to refer to datasets as such, instead of the older descriptor 'query'.
- We now use the inflector utility to pluralize groupings. Fixes "Querys", etc.
Trackster
-
Enhancements:
- Support large (10,000+) sets of chromosomes/contigs, as is often the case for low/non-model coverage genomes.
- Improve speed by streamlining data fetching and caching in Trackster.
- Add support for child tracks.
- UI hints for zooming out/in.
- Extend CIGAR string parsing to handle all operations (however, insertions still not fully supported).
- History dataset selection: show history name, show and order by hid in display.
- Modified interval index squish mode to display blocks and treat intervals as half-open.
-
Bug fixes:
- Fix CIGAR string parsing bug that prevented sequence data from being displayed correctly.
- BED files without score column or with scores that are floats can now be displayed.
- Better handling of error messages so that messages and data do not overlap.
User Interface (UI)
-
Search improvements:
- Size of search box increases to show help text (column label).
- Help text is highlighted rather than hidden when user clicks on search box.
- Standard and advanced search divs are toggled correctly when moving between searches.
Galaxy Reports
- Clarify in the reports config that Galaxy reports do not support SQLite.
- Add 2 new sample tracking reports and 2 new workflow reports, each of which provides the total number of items per month and the total number of items per user ( per month ).
Source
- Better egg version conflict resolution. Our eggs should always override dependencies installed elsewhere on
sys.path. - Job runners are now loaded dynamically; changes to the source are no longer required. Existing 3rd-party runners will need to add a class to instantiate to the runner-global 'all' list. See the provided runners for examples.
- The Galaxy-side code for the LWR job runner by John Chilton is now included. Documentation in the wiki is forthcoming, to learn more see John's documentation at: http://bitbucket.org/jmchilton/lwr/
- "LWR allows you to create a "cluster" out of any systems without installing specialized tightly integrated DRM software like PBS/SGE."
Bug Fixes
- Gracefully fall back to English if a client requests a locale not installed on the server.
- Don't attempt to check for the python-daemon egg on Python 2.4 (it's used by an optional experimental component and is not compatible with Python 2.4).
- The drmaa job runner would not always detect job completion on some systems. This change could also cause job failures to be considered successful job completions, but we have no way to test for these conditions. Please report if you discover this to be the case.
- Migrating to database version 62 no longer fails for MySQL. Thanks Leandro Hermida.
- Downloading eggs for offline systems works again.
About Galaxy
Galaxy is supported in part by NSF, NHGRI, the Huck Institutes of the Life Sciences, and The Institute for CyberScience at Penn State.